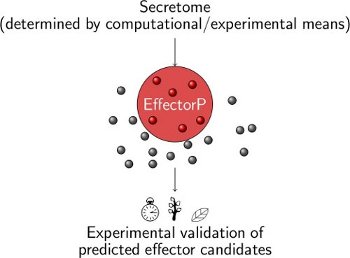
What is EffectorP?
Eukaryotic filamentous plant pathogens secrete effector proteins that modulate the host cell to facilitate infection. Computational effector candidate identification and subsequent functional characterization delivers valuable insights into plant-pathogen interactions. However, effector prediction in fungi has been challenging due to a lack of unifying sequence features such as conserved N-terminal sequence motifs. Fungal effectors are commonly predicted from secretomes based on criteria such as small size and cysteine-rich, which suffers from poor accuracy. Whilst in oomycetes sequence motif and Hidden Markov Model searches are well-established methods to predict some classes of cytoplasmic effectors, it will miss effectors that do not carry such motifs or domains as well as apoplastic effectors.
EffectorP is a machine learning method for fungal and oomycete effector prediction in secretomes and has been trained to distinguish secreted proteins
from secreted effectors. EffectorP does not use sequence motif searches (such as RxLR) for effector prediction in oomycetes.
EffectorP improves fungal effector prediction from secretomes based on a robust signal of sequence-derived properties, achieving accuracy of over 85%. For more details on the underlying methods, please see our papers:
Sperschneider J and Dodds PN (2021)
EffectorP 3.0: prediction of apoplastic and cytoplasmic effectors in fungi and oomycetes.
MPMI.
Abstract
Sperschneider J, Dodds PN, Gardiner DM, Singh KB, Taylor JM (2018)
Improved prediction of fungal effector proteins from secretomes with EffectorP 2.0.
Molecular Plant Pathology.
Abstract
Sperschneider J, Gardiner DM, Dodds PN, Tini F, Covarelli L, Singh KB, Manners JM, Taylor JM (2015)
EffectorP: Predicting Fungal Effector Proteins from Secretomes Using Machine Learning.
New Phytologist, doi:10.1111/nph.13794.
Abstract
EffectorP improves fungal effector prediction from secretomes based on a robust signal of sequence-derived properties, achieving accuracy of over 85%. For more details on the underlying methods, please see our papers:
Input
EffectorP has been trained to find fungal or oomycete effectors in secretomes, so please submit a FASTA file of secreted proteins to test
if they are predicted effectors. It is recommended to use tools such as SignalP or
Phobius to predict first if a protein is likely to be secreted.
Alternatively, experimentally determined secretomes instead of computationally predicted secretomes
may be submitted to EffectorP.
Do not use EffectorP to scan the whole protein set of a fungal or oomycete genome.
EffectorP has to be run on a secretome, as it recognizes a cytoplasmic signal also in intracellular, non-secreted proteins.